-

Numbat: haplotype-aware CNV analysis from scRNA-seq
Numbat is a haplotype-enhanced CNV caller for scRNA-seq data. It integrates signals from gene expression, allelic ratio, and population-derived haplotype information to accurately infer allele-specific CNVs in single cells and reconstruct their lineage relationship. Please see the paper and GitHub repository for details on the method.
-
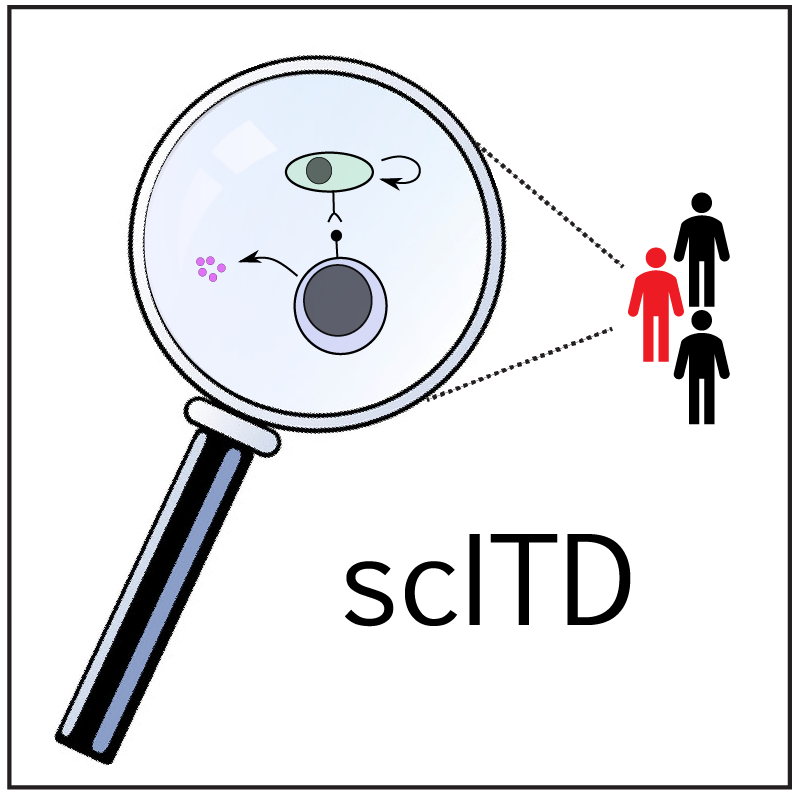
scITD: single-cell interpretable tensor decomposition
scITD (Single-Cell Interpretable Tensor Decomposition) is a computational method capable of extracting multicellular gene expression programs that vary across donors or samples. The approach is premised on the idea that complex biological processes often involve the coordinated actions and interactions of multiple cell types. Given single-cell expression data from multiple heterogeneous samples, scITD aims to infer these joint patterns of dysregulation impacting multiple cell types. The preprint and GitHub repository are also available.
-
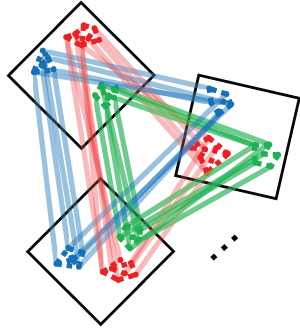
Conos: wiring together scRNA-seq dataset collections
Conos (Clustering on Network of Samples) is a tool for the joint analysis of heterogeneous collections of scRNA-seq datasets, such as collections combining multiple individuals, conditions, tissues, or technological platforms. Please see the publication for a detailed description and analysis examples, as well as the GitHub repository for hands-on tutorials and source code.
-

RNA velocity estimation
velocyto is a framework which predicts the movement of cells in transcriptional space, by estimating the first derivative of the transcriptional state - RNA velocity. This provides a basis for quantitative modeling of dynamic biological processes, such as cell differentiation, or perturbation response.
-
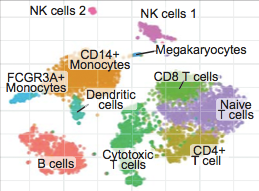
Demultiplexing of single-cell RNA-seq data
dropEst is a pipeline for demultiplexing single-cell RNA-seq data, implementing additional corrections for accurate estimation of the molecular count matrices. Please refer to the original publication for details.
-

Single Cell Transcriptional Analysis
We have developed the R package pagoda2 for analyzing and interactively exploring large-scale single-cell RNA-seq datasets. Please see the GitHub repository for hands-on tutorials and source code. The tutorial can also be found here.
The SCDE package provides routines for analysis of single-cell RNA-seq data. It is based on the probabilistic mixture error model, which is used to implement differential expression, subpopulation analysis and other tasks on the data.
-
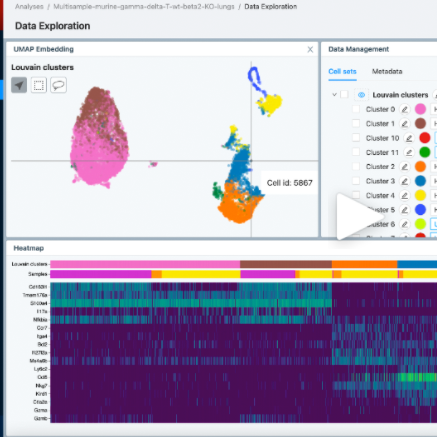
Cellenics
Cellenics is a user-friendly online tool for single cell RNA-seq data analysis (currently only supports 10X Chromium datasets). The platform is designed specifically for biologists. It provides automatic data processing, pre-loaded interactive plots, a point-and-click interface to fully explore your data, together with the option to customize and export publishable-quality figures.
-

Transposable Element Analyzer
The Tea pipeline is designed to identify insertions of repetitive elements (such as LINE1 repeats or endogenous retroviral elements) in the human genomes. Its primary aim is to detect novel repeat insertions occurring in somatic tissues (e.g. cancerous tumors), however it is also capable of detecting instances of repeat insertions polymorphic among individuals.
For more details, please refer to the Tea manuscript, and the pipeline download page on the Park Lab site
-
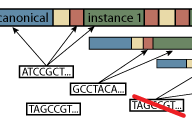
Repeat Enrichment Estimator
The software, developed during PI's fellowship in the Park Lab, provides means to estimate the enrichment of repetitive elements in the short-read sequencing data. For details, please refer to the manuscript. The implementation, custom sequence assemblies are available from the Park Lab server, which also provides a web interface for running the analysis.
-
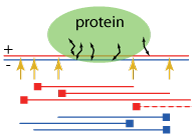
ChIP-seq processing pipeline (SPP)
The spp R package provides routines for processing ChIP-seq data. It supports the output of many short-read aligners, and can be used to determine a statistically significant set of binding peaks or broad regions of enrichment. The details of peak-calling algorithms were described in the initial manuscript. The updated versions of the package, and a brief tutorial are available from the Park Lab site.
Kharchenko Lab
Making sense of genomic data!
 National Science Foundation
National Science Foundation  National Institutes of Health
National Institutes of Health